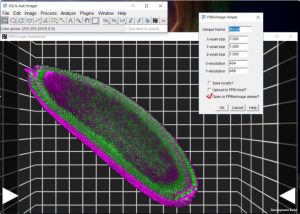
A yellow line should appear on stacked image To measure fluorescent intensity: draw line (or other shape chosen from Window appears with Colour and possibility to tick boxes of both channels.įirst tick box of Channel 1 and click More to assign appropriate colourĬhange Colour to Composite to view stack with both colours. 
To question asking: Convert to multi-channelĭisplay mode: change from Composite to Colour and click OK To assign the right colour to each channel, click on "Image => “Colour” =>Ĭhange Composite to Colour. Right to switch between channels both channels are shown in the same Now both channels appear in the same window (you need to scroll left and Using ImageJ to Measure Intensity of FluorescenceĢ.Go to "Image => "Stacks => “Images to Stack” Name stack, e.g. I have been provided the following protocol: What information are you interested in getting from this image? They are breast cancer cell lines called BT549 What is the image about? Provide some background and/or a description of the image. My task is to determine if the glycosidase inhibitor treatment that I am asked to evaluate has indeed altered the cellular glycosylation by examining the lectin florescence stating of the cancer cells for those treated with my compound and compared to the untreated and the positive control (deoxynojirimycin). Scan_Plate_R_p00_0_B02f00d1.TIF is the green image Analysis goals Scan_Plate_R_p00_0_B02f00d0.TIF is blue image I have been told to use the * “scan _Plate_R…” for my analysis. Images labelled “scan _Plate_R…” split the green (lectin) and blue(DAPI, nuclear stain). Have green (lectin) and blue (DAPI, nuclear stain) overlaid (merged) Share a minimal working example of your macro code.Upload an original image file here directly or share via a link to a file-sharing site (such as Dropbox) – (make sure however that you are allowed to share the image data publicly under the conditions of this forum).For other applications, I’ve seen people additionally convert their images in ImageJ into 32-bit or 8-bit formats prior to analyzing them. tif format for the analysis in order to prevent compression. oif format, and I was going to export them into a. Q3: How do I control for background fluorescence? Is there a way to “invert” the threshold that I set to measure the integrated density of the area surrounding the fluorescent protein so that I can subtract it from the integrated density of the fluorescent protein? Q2: If I manually threshold the image, do I need to make sure that I use the exact same settings for each image to eliminate any bias in thresholding? If I do, I’m assuming these settings would be based upon a control, such as wild-type, untreated cells. Will this help me to get the measurements that I need? In brief, I was going to duplicate the image, manually threshold to select only the green fluorescence, perform a watershed separation, and then analyze the particles to measure the integrated density, making sure to redirect the threshold I set to the original image. Q1: I was planning on using the steps outlined on the “Particle Analysis” page of the ImageJ cookbook to measure the integrated density. I’d like to measure the fluorescence intensity of the entire image and then divide it by the number of cells in the image (which I will manually count) in order to get an average intensity per cell. There are around 20-30 cells in each image. I’ve included a sample image below of the fluorescence I’m hoping to measure. I’ve read several ImageJ articles and forum posts about this topic, but I’m having difficulty connecting what I’ve read to create an analysis protocol for my experiment. Hi everyone! I’m a new user of ImageJ, and I’m interested in measuring and comparing the fluorescence intensity of antibody-stained cells across a variety of conditions (wild-type vs.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
